LipidFinder: A computational workflow discovery of lipids
LipidFinder is an open-source Python workflow which searches a number of different databases to obtain putative identification of lipids, and assigns them to a class based on the LIPID MAPS® classification system. The software quickly distinguishes and quantifies lipid-like features from contaminants, adducts and noise in high resolution liquid chromatography/mass spectrometry (LC/MS) datasets that have been pre-aligned using XCMS.
Note that your data needs to be high resolution MS (e.g. at least 60,000ppm) and with long chromatography to separate isobaric lipids. The approach is not suitable for shotgun lipidomics, MS/MS or low resolution datasets. Data provided in our demo section used an Orbitrap Elite Mass Spectrometer, but the software is MS platform independent.
We recommend the user to read the detailed about LipidFinder before using the first time. Additionally, we have performed a comparison analysis between XCMS and XCMS + LipidFinder. You can see the results .
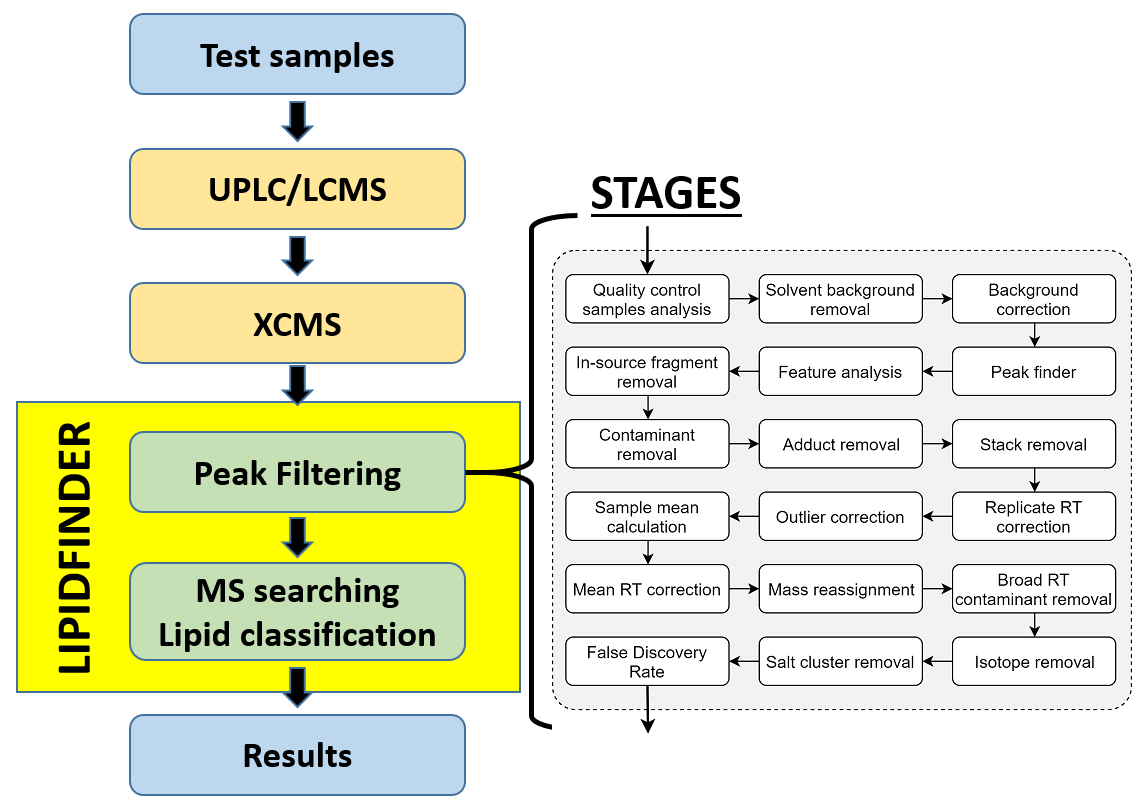
 LipidFinder: A computational workflow for discovery of lipids identifies eicosanoid-phosphoinositides in platelets.
A. O'Connor, C.J. Brasher, D.A. Slatter, S.W. Meckelmann, J.I. Hawksworth, S.M. Allen and V.B. O'Donnell.
JCI Insight. 2017;2(7):e91634. doi:10.1172/jci.insight.91634
LipidFinder on LIPID MAPS®: peak filtering, MS searching and statistical analysis for lipidomics.
Fahy E, Alvarez-Jarreta J, Brasher CJ, Nguyen A, Hawksworth JI, Rodrigues P, Meckelmann S, Allen SM, O'Donnell VB.
Bioinformatics. 2019 Feb 15;35(4):685-687. doi: 10.1093/bioinformatics/bty679.
LipidFinder 2.0: advanced informatics pipeline for lipidomics discovery applications.
J. Alvarez-Jarreta, P.R.S. Rodrigues, E. Fahy, A. O’Connor, A. Price, C. Gaud, S. Andrews, P. Benton, G. Siuzdak, J.I. Hawksworth, M. Valdivia-Garcia, S.M. Allen, V.B. O’Donnell
LipidFinder: A computational workflow for discovery of lipids identifies eicosanoid-phosphoinositides in platelets.
A. O'Connor, C.J. Brasher, D.A. Slatter, S.W. Meckelmann, J.I. Hawksworth, S.M. Allen and V.B. O'Donnell.
JCI Insight. 2017;2(7):e91634. doi:10.1172/jci.insight.91634
LipidFinder on LIPID MAPS®: peak filtering, MS searching and statistical analysis for lipidomics.
Fahy E, Alvarez-Jarreta J, Brasher CJ, Nguyen A, Hawksworth JI, Rodrigues P, Meckelmann S, Allen SM, O'Donnell VB.
Bioinformatics. 2019 Feb 15;35(4):685-687. doi: 10.1093/bioinformatics/bty679.
LipidFinder 2.0: advanced informatics pipeline for lipidomics discovery applications.
J. Alvarez-Jarreta, P.R.S. Rodrigues, E. Fahy, A. O’Connor, A. Price, C. Gaud, S. Andrews, P. Benton, G. Siuzdak, J.I. Hawksworth, M. Valdivia-Garcia, S.M. Allen, V.B. O’DonnellBioinformatics. 2020 October 07. doi: 10.1093/bioinformatics/btaa856 LipidFinder's source code for workstations is available on GitHub (https://github.com/ODonnell-Lipidomics/LipidFinder), which includes an amalgamator to work with both ion polarities, multiple database classification and user manuals. If you have any comments or suggestions, please send an e-mail to: lipidfinder@cardiff.ac.uk